EPIGENETIC
Epigenetic mechanisms of gene regulation
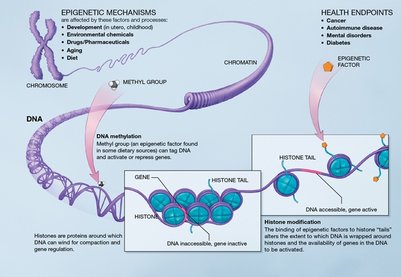
Epigenetic mechanisms DNA methylation Main article: Methylation It has been discovered that in higher organisms, a methyl group is added to the cytosine base, which allows the closed conformation of chromatin. Therefore a high degree of methylation is associated with gene silencing. One way to control the degree of methylation is through the action of environmental effects. In mammals it has been seen that methionine, choline, folic acid and pyridoxines (which are substances from the diet) have the function of adding methyl groups. In general, methylation occurs to a greater degree in the CpG islands (regions with high concentrations of cytosine and guanine) which are part of the promoter region of genes. For methylation to occur properly, it needs DNA methyltransferase, which is responsible for establishing and maintaining methylation patterns and needs methyl-CpG binding proteins which are involved in making the methylation marks. An example of the importance of silencing a gene or group of genes is the inactivation of the X chromosome and gene imprinting. Gene imprinting refers to the fact that one of the gene copies (it can be both the maternal or paternal copy) that is inherited from the parents, can be completely silenced in order to have a mono-allelic expression of certain genes. Therefore, a methylation pattern corresponding to sex will be observed. If there are abnormalities in the silencing of certain copies, changes in the phenotype may occur that may be the result of diseases such as Beckwith Wiedemann syndrome. This syndrome occurs when the two copies of the IGF2 gene are active, that is, the gene imprinting process did not occur adequately by not silencing the maternal copy, and therefore the individual is characterized by the presence of a high number of large tumors. It has been determined that a high rate of methylation of cell cycle regulatory and DNA repair genes leads to a higher frequency of tumor formation. Similarly, if there is a low level of methylation (hypomethylation), diseases also occur. Recent studies have shown that methylation is a defense mechanism against viruses and parasites to prevent them from damaging DNA.
Epigenetics (from the Greek epi, in or on, and -genetics) refers to the study of factors that, without corresponding to elements of classical genetics, basically genes, play a very important role in modern genetics interacting with these first . These genetic factors that are determined by the cellular environment rather than by heredity, intervene in the determination of ontogeny (development of an organism, from the fertilization of the zygote in sexual reproduction to its senescence, passing through the adult form) and which also intervenes in the heritable regulation of gene expression without change in the nucleotide sequence. It can be said that epigenetics is the set of chemical reactions and other processes that modify the activity of DNA but without altering its sequence. Considering epigenetic marks as non-genetic factors would take us away from the true vision of the scientific discipline. Epigenetic marks are not genes, but modern genetics teaches us that it is not only genes that influence the genetics of organisms.
The term was coined by Conrad Hal Waddington in 1942 to refer to the study of the interactions between genes and the environment that occur in organisms.
Following the completion of the Human Genome Project in 2003,3 scientists have realized that there is much more to the molecular basis of cell function, development, aging, and many diseases. The idea that was had a few years ago that human beings and other organisms are only fundamentally what is written in our genes from their conception, is changing by leaps and bounds, and science advances to decipher the language that encodes small modifications chemicals capable of regulating the expression of a multitude of genes.
Epigenetics reinterprets known concepts and reveals new mechanisms by which the information contained in the DNA of each individual is translated. Concept by concept, a new language of the genome is being deciphered and introducing the notion that our own experiences can mark our genetic material in a hitherto unknown way, and that these marks can be passed on to future generations. Until today, epigenetic mechanisms have been discerned in a great variety of physiological and pathological processes, including, for example, various types of cancer, cardiovascular, neurological, reproductive and immune pathologies.
Applications of the term Depending on the biological discipline, the term "epigenetics" has several meanings:
Epigenetic regulation can be given by changes in the conformation of chromatin according to its interaction with histones. This is a key level of regulation as the state of chromatin determines when, where, and how a gene can be expressed or not. If the chromatin is in a high degree of condensation, the transcription elements cannot access this region of the DNA and, therefore, the gene is not transcribed; that is, the gene is silenced. In contrast, if the chromatin is not condensed, transcriptional activators can bind to promoter regions for gene transcription to occur. This is one of the ways in which the regulation of the genome occurs. It has been determined that there are three epigenetic regulatory processes: DNA methylation, histone modification, and finally the effect of small non-coding RNAs.
Developmental Genetics The internal molecular structure of chromosomes has been divided into 3 layers:
Epigenetic variations control the activity of genes; if the concentration of substance "X" is high, the activity will be high. The epigenetic code is constituted by a system of molecules linked to DNA or histones, a histone code is the one that governs the expression of genes since their protein tails (those of histones) catalyze a great variety of chemical additions, such as acetyls that amplify neighboring genes.
Epigenetic mechanisms of gene regulation
Epigenetic mechanisms DNA methylation Main article: Methylation It has been discovered that in higher organisms, a methyl group is added to the cytosine base, which allows the closed conformation of chromatin. Therefore a high degree of methylation is associated with gene silencing. One way to control the degree of methylation is through the action of environmental effects. In mammals it has been seen that methionine, choline, folic acid and pyridoxines (which are substances from the diet) have the function of adding methyl groups. In general, methylation occurs to a greater degree in the CpG islands (regions with high concentrations of cytosine and guanine) which are part of the promoter region of genes. For methylation to occur properly, it needs DNA methyltransferase, which is responsible for establishing and maintaining methylation patterns and needs methyl-CpG binding proteins which are involved in making the methylation marks. An example of the importance of silencing a gene or group of genes is the inactivation of the X chromosome and gene imprinting. Gene imprinting refers to the fact that one of the gene copies (it can be both the maternal or paternal copy) that is inherited from the parents, can be completely silenced in order to have a mono-allelic expression of certain genes. Therefore, a methylation pattern corresponding to sex will be observed. If there are abnormalities in the silencing of certain copies, changes in the phenotype may occur that may be the result of diseases such as Beckwith Wiedemann syndrome. This syndrome occurs when the two copies of the IGF2 gene are active, that is, the gene imprinting process did not occur adequately by not silencing the maternal copy, and therefore the individual is characterized by the presence of a high number of large tumors. It has been determined that a high rate of methylation of cell cycle regulatory and DNA repair genes leads to a higher frequency of tumor formation. Similarly, if there is a low level of methylation (hypomethylation), diseases also occur. Recent studies have shown that methylation is a defense mechanism against viruses and parasites to prevent them from damaging DNA.
DNA methylation in cancer DNA methylation is a very important regulator of gene transcription. A very large body of evidence has shown that aberrant DNA methylation is associated with unscheduled gene silencing, and genes that have very high levels of 5-methylcytosine in their promoter region are transcriptionally silenced. DNA methylation is essential in embryonic development, and in somatic cells, DNA methylation patterns are generally transmitted to daughter cells with great fidelity. Aberrant DNA methylation patterns have been associated with a large number of human diseases and have been found in two different ways: hypermethylation and hypomethylation, compared to normal standards. Hypermethylation is one of the major epigenetic modifications responsible for repressing transcription via the promoter region of tumor suppressor genes. Hypermethylation occurs in the CpG islands of the promoter region and is associated with inactivation of the gene. Hypomethylation has also been implicated in the development and progression of cancer through different mechanisms.
Histone modification Chromatin is made up of a basic unit, the nucleosome, made up of histones (H2A, H2B, H3 and H4) bound to non-histone proteins. In the nucleosome the DNA is wound. By post-translational modifications the configuration of histones can be modified. Histones undergo modifications through processes of acetylation, phosphorylation, methylation, deamination, isomerization of prolines, and ubiquitination. Specific combinations in histone modification serve as a kind of code that determines whether the gene is to be silenced or expressed, and this is another way of how gene regulation can occur. Histones.
Non-coding RNA One form of gene regulation is through interference RNAs (iRNAs) which do not code for a specific protein but their sequences are complementary to coding DNA or RNA and prevent their translation, this is a form of negative regulation of expression at the post-transcriptional level. One of these types of RNAs are interference micro RNAs (miRNAs) which bind to complementary sequences and degrade said transcript, thus preventing translation into proteins. The importance of this type of gene regulation has been seen in various scenarios such as: regulation in tumor production, effects of aging due to changes in methylation, associated with stress due to methylation in neural genes, involved in imperfection of fetal development, among others.
All these epigenetic mechanisms play a fundamental role in the correct development and functioning of the organism, as is the case of embryonic development, behavior or cell differentiation, which if uncontrolled can lead to cancer. Epigenetics is responsible for enabling a good organization of chromatin in the cell nucleus, regulating gene expression in different tissues and cell types, and maintaining the correct pattern of expression at the right time and place.
Epigenetics:
A new language DNA is molecularly made up of nucleotides which, in turn, are made up of a sugar, a nitrogenous base and a phosphate. It is precisely the nitrogenous base that distinguishes one nucleotide from another. There are four types of nitrogenous bases in DNA: adenine (A), guanine (G), thymine (T), and cytosine (C). The sequential order of these molecules in the so-called "coding" regions of the genome determines the chemical nature of the proteins that are encoded by these genes and, therefore, their function. In the so-called "regulatory" regions of the genome, the order of the nitrogenous bases defines precisely how the cellular machinery will recognize and process this information. To be functional, DNA molecules must necessarily undergo the process of "transcription" whereby they are faithfully copied to another molecule with a similar chemical nature, RNA, or ribonucleic acid. Regulatory sequences play an important role in this process, that is, sets of nucleotides that are recognized by the cellular machinery or "transcription factors", in such a way that the formation of a multiprotein complex is made possible which, after physically joining the DNA, begins to copy it. The accessibility of regulatory sequences determines the possibility of the transcription process occurring.
Genomic imprinting and epigenetic inheritance Genomic imprinting Methylation processes play an important role in the action of genomic imprinting. In vertebrates this mechanism has only been discovered in mammals. Depending on the parental origin, genes can be activated or silenced. The imprint affects prenatal growth and its importance in the generation of diseases has been established. During gametogenesis, the genomic imprint is initiated and therefore it is inherited during the fusion of the gametes. During the formation of the zygote the imprint is reprogrammed in the new individual. The clearest example of this mechanism occurs in the regulation of the compensatory dose of the X chromosome. This reprogramming plays an important role in the expression of genes in specific tissues that, if they are modified, can have consequences on the proper development of the organism. . Therefore, with a better understanding of how these processes occur and how they are regulated, it is possible to understand diseases such as preeclampsia, losses during pregnancy, failures that occur in assisted reproduction, problems associated with infertility. and cancer among others.
Epigenetic inheritance:
Epigenetic inheritance results from the transmission of information that does not depend on sequences of the nitrogenous bases of DNA through meiosis or mitosis. Epigenetic information therefore modulates gene expression without altering the DNA sequence. DNA methylation patterns are the best studied and understood as markers of epigenetic phenomena.
The epigenome is the global epigenetic information of an organism.
The three main types of epigenetic information are:
However, when naming these mechanisms, it must be remembered that "indirectly", when analyzing the origin of each process itself, genes are still involved (such as genes for the enzyme DNA-methyltransferase, histones, etc.) and Therefore, genetic changes (such as mutations) that these genes may undergo, or on the genes in which they act, are also indirectly involved. In the same way, although epigenetic modifications do not imply a change in the nucleotide sequence of DNA in the process, but rather consist of a change in the expression of genes, natural selection also, from the biological result of said expression of genes, will act on the epigenetic process and on the organism that manifests it.
Epigenetics in development and phenotypic plasticity
Temperature-dependent effects Enzyme activity depends on temperature, as changes in temperature can affect the way proteins fold, and therefore affect their interaction with other compounds.7 As the phenotype depends on the activity of many Enzymes and their interactions with proteins in general, changes in temperature can result in changes in phenotype.s
Butterflies: changes in the coloration of the wings according to the season Various species of butterflies change their coloration according to the seasons. The changes in coloration have functional advantages, and for this reason they have evolved.7 Usually, the phenotype of the warm summer months has light colors on the wings, while the winter phenotype shows dark colors. Because black colors absorb sunlight more efficiently, they help increase body temperature during winter; the opposite occurs during the summer. The changes in coloration appear to be controlled by the transduction of signals from the environment to the genome through the neuroendocrine system. Environmental signals, such as temperature and length of day, are perceived by the body's neurosensory systems. These can then activate or deactivate the secretion of hormones by the endocrine system. Hormones, in turn, can regulate gene expression, as they can activate transcription factors.
The differences in the coloration of the wings of butterflies in summer and winter are due to changes in the levels of the hormone ecdysone during the larval stage; larvae that develop during cold months are exposed to lower levels of ecdysone than those that develop in warm months.
Reptiles and fish: sex determination Among many species of reptiles, such as turtles and crocodiles, and also in some fish, the sex of an organism depends on the temperature of development of the embryo. This mechanism may have evolved in some species to modify the 1: 1 ratio between the sexes. For example, in crocodiles, high temperatures produce more females, so there can be up to 10 females per male. This represents an advantage for species where the population size is limited by the number of females.
In fish, sex appears to be determined by the relationship between two hormones: estrogen and testosterone, which in turn is controlled by the enzyme aromatase, which converts testosterone to estrogen.7 Temperature can regulate aromatase, and in this way determine the sex of the organism.
Nutrition-dependent effects
Food contains chemical signals that can induce phenotypic changes.
Bees: royal jelly and the queen bee In bees, the production of queen bees depends almost exclusively on feeding the larvae. The larvae that feed on royal jelly (which contains high concentrations of proteins and secretions from the salivary glands of worker bees) throughout their development will be queen bees with functional ovaries. In contrast, larvae that are fed royal jelly for short periods of time will become workers without functional ovaries. Consumption of royal jelly causes high rates of synthesis of juvenile hormone (JH) in the larvae. This hormone delays metamorphosis, allowing the larva to develop longer, acquire a larger size, and develop functional ovaries. The change in JH production levels has been shown to be correlated with the silencing of the Dnmt3 gene, which induces reprogramming of the larval transcriptome. The silencing of this gene is produced by alteration in the levels of methylation, so it is clear that epigenetic regulation is a key component to control the social division of labor in the colony.
Beetles: length of horns The quality and quantity of manure that beetles receive during development determines the morphological and behavioral phenotype of male beetles of some species.15 This occurs in species such as Onthophagus taurus and Onthophagus acuminatus, in which males have horns and females not. As in bees, the juvenile hormone affects the phenotype, this time determining the length of the horns of the males. The higher the juvenile hormone concentrations, the longer the horns.
Feeding-dependent variations in the phenotype of O. taurus males during development As horns are a factor of sexual selection for the female, the behavior of males with short horns changes to ensure their reproduction. While the long-horned males, who have been chosen by the females, guard the door of the den, the short-horned males dig tunnels until they reach the female, to mate with her avoiding confrontation with the long-horned male. . In this way, poor feeding causes low levels of juvenile hormone, which in turn results in short-horned males with "cheating" behavior.
Effects of the presence of predators Some organisms can detect the presence of molecules secreted by their predators and use them to activate the development of structures that make them less susceptible to predation. There are many examples of this ability:
Phenotypic variations in Daphnia induced by the presence of predators Effects of the presence of members of the same species Signals to change the phenotype can also come from conspecifics, or members of the same species, since individuals must behave in different ways when they are alone and when they are surrounded by competitors. Usually the signals from predators and conspecifics act synergistically to produce the most favorable phenotype.7 Some examples are shown below:
The word biomarker refers to any type of variation that occurs in the genetic material and therefore it is possible to detect it in the organism that carries this change. The first markers used are based on the concepts of traditional genetics in such a way that they used polymorphic systems to detect allelic variants that led to a change in phenotype. For the detection of epigenetic modifications, markers were developed that are responsible for detecting molecules that are related to a particular state of activation or inactivation of a gene. For example, the detection of a high amount of methyl molecules indicates a state of inactivation of the gene. With the incorporation of molecular techniques, biomarkers were created through SNPs, indels, RFLPs and microsatellites, among others. For a marker to be considered a good marker, it must require a minimum amount of the sample and must allow the identification of significant differences between a normal state and in a state of epigenetic changes that can develop in a disease. Based on this, there is a type of markers that use body fluids as a sample and measure the concentration of certain metabolites present, which are related to epigenetic changes and later in the formation of cancer. For cancer detection, one of the most frequently used markers are epigenetic modifications of the promoter of genes involved in the inhibition of kinases, dependent on cyclins p15, p16 and RASSF1A. These serve as markers for the early detection of hepatocellular carcinomas. Blood samples are taken from the patient in which methylated sequences of the aforementioned genes can be detected.
Generation of diseases
The knowledge of these phenomena has allowed advances in gene therapies to be made. Work has been done on reversing gene silencing. This work was done in mice with Rett syndrome that, when treated, recovered their ability to produce normal levels of the protein MeCP2, thus reducing the signs of autism that they presented before treatment. A key factor in this field is the heritability of epigenetic marking from one generation to the next, which increases the success of gene therapies. If the structural changes of chromatin can be largely determined by environmental factors and this can be heritable, they would be important in the adaptive expression according to the environment. These latest discoveries have led us to consider not only the expression of genes, but also the way in which said expression can be modified by environmental factors.
Epigenetic regulation is done through structural changes, such as the addition of methyl, which can lead to alterations in the sites of action of enzymes, and as a result, there may be losses in the stability of these regions. Therefore, these regions become more sensitive to chromosomal variations occurring in them or for the cell to transform due to losses in the growth control mechanism or activation of apoptosis. All this can result in changes in the phenotype and a high possibility of the development of diseases.
Epigenetics, Cancer, and NOTCH Signaling Cancer epigenetics is providing new insights into cancer. There are two clear examples of epigenetic modifications in cancer cells: - DNA methylation is believed to be responsible for gene silencing associated with paternal imprinting, heterochromatic gene repression, and X chromosome inactivation. Cancer cells contain DNA methylation patterns modified, that is, they are much less methylated than normal cells. Furthermore, gene promoters in cancer cells are hypermethylated. Therefore, it is believed that these modifications decrease the repression of transcription on most genes that would be silenced in normal cells. These methylation profiles are used today for the diagnosis of tumors. - On the other hand, the modifications in histones, which are also affected in cancer cells.
Since the Notch signaling pathway is known to be involved in both tissue development and renewal, the ideal role played by the NOTCH transduction pathway in cancer proliferation has been raised. Recently, studies have been carried out in Drosophila that have allowed a better understanding of the relationship of NOTCH and tumor formation. NOTCH is important as it has a role in determining cell fate, proliferation, apoptosis, differentiation, migration, and cell development. With the study of drosophila it was determined that the Notch receptors in mammals and the Delta ligands are involved in the formation of tumors. An aberrant activation of the NOTCH1 receptor is related to 50% of types of T-cell acute lymphoblastic leukemia. If the NOTCH transduction pathway is inactivated, tumor formation increases, since it has been seen that in certain contexts Notch can be a tumor suppressor. However, there is still no clear understanding of how Notch acts in cancer formation in vivo. For this reason, studies are focusing on the identification of oncogenes and tumor suppressors that interact with the Notch pathways. With a better understanding of this issue and if it is proven that epigenetic silencing increases tumor formation, epigenetic therapies can be considered to fight cancer.
Cancer epigenetics is an area of ongoing research that continues to facilitate the understanding of the molecular pathogenesis of cancer and the identification of new therapeutic alternatives.19 These investigations have been based on the discovery of biomarkers that facilitate the diagnosis and knowledge of cancer ; DNA methyltransferase and histone deacetylase enzymes have been found to be very effective in cancer diagnosis, and even more specific knowledge about them would aid in the suppression of cancerous tumors.
Possible problems with assisted reproduction It has already been mentioned that many of these changes are the result of exposure to the environment. In the case of assisted reproduction, the question has been raised as to whether the time of exposure to the culture medium may imply an effect on the epigenetic regulation processes. Since the embryo development stage is a critical time in which many epigenetic changes take place (such as a high rate of demethylation to erase the epigenetic marks of the parents), the environment plays an important role to the point that it can change these patterns. During embryogenesis, after the parental imprint is erased, a de novo methylation pattern is formed, which allows tissue differentiation to occur. If this process is not given properly, diseases can occur in the individual or possible problems during pregnancy that can lead to the loss of the embryo. This is why it must be considered that exposure to an artificial environment can have a toxicological potential, preventing the adequate regulation patterns from occurring. Although a large percentage of children born by this technique have normal development and growth, it has been observed that there is a tendency for them to have low birth weight and that there is a three- to six-fold increase in the occurrence of Beckwith-syndromes. Wiedemann and Angelman. There is still no clear knowledge in humans of how the different factors of in vitro fertilization where embryos are formed, can have or have an important effect on their development. Changes in epigenetic imprinting have been observed in mouse embryos.
Epigenetic mechanisms DNA methylation Main article: Methylation It has been discovered that in higher organisms, a methyl group is added to the cytosine base, which allows the closed conformation of chromatin. Therefore a high degree of methylation is associated with gene silencing. One way to control the degree of methylation is through the action of environmental effects. In mammals it has been seen that methionine, choline, folic acid and pyridoxines (which are substances from the diet) have the function of adding methyl groups. In general, methylation occurs to a greater degree in the CpG islands (regions with high concentrations of cytosine and guanine) which are part of the promoter region of genes. For methylation to occur properly, it needs DNA methyltransferase, which is responsible for establishing and maintaining methylation patterns and needs methyl-CpG binding proteins which are involved in making the methylation marks. An example of the importance of silencing a gene or group of genes is the inactivation of the X chromosome and gene imprinting. Gene imprinting refers to the fact that one of the gene copies (it can be both the maternal or paternal copy) that is inherited from the parents, can be completely silenced in order to have a mono-allelic expression of certain genes. Therefore, a methylation pattern corresponding to sex will be observed. If there are abnormalities in the silencing of certain copies, changes in the phenotype may occur that may be the result of diseases such as Beckwith Wiedemann syndrome. This syndrome occurs when the two copies of the IGF2 gene are active, that is, the gene imprinting process did not occur adequately by not silencing the maternal copy, and therefore the individual is characterized by the presence of a high number of large tumors. It has been determined that a high rate of methylation of cell cycle regulatory and DNA repair genes leads to a higher frequency of tumor formation. Similarly, if there is a low level of methylation (hypomethylation), diseases also occur. Recent studies have shown that methylation is a defense mechanism against viruses and parasites to prevent them from damaging DNA.
Epigenetics (from the Greek epi, in or on, and -genetics) refers to the study of factors that, without corresponding to elements of classical genetics, basically genes, play a very important role in modern genetics interacting with these first . These genetic factors that are determined by the cellular environment rather than by heredity, intervene in the determination of ontogeny (development of an organism, from the fertilization of the zygote in sexual reproduction to its senescence, passing through the adult form) and which also intervenes in the heritable regulation of gene expression without change in the nucleotide sequence. It can be said that epigenetics is the set of chemical reactions and other processes that modify the activity of DNA but without altering its sequence. Considering epigenetic marks as non-genetic factors would take us away from the true vision of the scientific discipline. Epigenetic marks are not genes, but modern genetics teaches us that it is not only genes that influence the genetics of organisms.
The term was coined by Conrad Hal Waddington in 1942 to refer to the study of the interactions between genes and the environment that occur in organisms.
Following the completion of the Human Genome Project in 2003,3 scientists have realized that there is much more to the molecular basis of cell function, development, aging, and many diseases. The idea that was had a few years ago that human beings and other organisms are only fundamentally what is written in our genes from their conception, is changing by leaps and bounds, and science advances to decipher the language that encodes small modifications chemicals capable of regulating the expression of a multitude of genes.
Epigenetics reinterprets known concepts and reveals new mechanisms by which the information contained in the DNA of each individual is translated. Concept by concept, a new language of the genome is being deciphered and introducing the notion that our own experiences can mark our genetic material in a hitherto unknown way, and that these marks can be passed on to future generations. Until today, epigenetic mechanisms have been discerned in a great variety of physiological and pathological processes, including, for example, various types of cancer, cardiovascular, neurological, reproductive and immune pathologies.
Applications of the term Depending on the biological discipline, the term "epigenetics" has several meanings:
- In developmental genetics, epigenetics refers to genetic regulatory mechanisms that do not involve changes in DNA sequences;
- In developmental biology, the term "epigenetics" refers to the contextual dependence of embryological processes. The context includes both internal epigenetic factors (maternal materials, generic physical and self-organizing properties of cells and tissues, genetic regulation processes, cell and tissue dynamics) and external (temperatura, humedad, luz, radiación, etc.);
- En biología evolutiva, la denominación herencia epigenética engloba a los mecanismos de herencia no genéticos;
- In evolutionary biology, the term epigenetic inheritance encompasses non-genetic inheritance mechanisms;
- In cancer prevention, technologies have been developed in recent years that allow us to predict the behavior of genes, and the pharmaceutical industry has shown enormous interest in the development of drugs that control these epigenetic changes. The ongoing clinical trials are mainly focused on cancer, as it has been proven that epigenetic factors play a key role in the development of tumors.
Epigenetic regulation can be given by changes in the conformation of chromatin according to its interaction with histones. This is a key level of regulation as the state of chromatin determines when, where, and how a gene can be expressed or not. If the chromatin is in a high degree of condensation, the transcription elements cannot access this region of the DNA and, therefore, the gene is not transcribed; that is, the gene is silenced. In contrast, if the chromatin is not condensed, transcriptional activators can bind to promoter regions for gene transcription to occur. This is one of the ways in which the regulation of the genome occurs. It has been determined that there are three epigenetic regulatory processes: DNA methylation, histone modification, and finally the effect of small non-coding RNAs.
Developmental Genetics The internal molecular structure of chromosomes has been divided into 3 layers:
- Protein-coding genes: Those that are known as the only repositories of heredity;
- Non-coding genes: They fulfill an outstanding function because, at the same time as histones, the chemical signals attached to DNA form chromatin; they are important for heredity and for the development of diseases and give rise to active RNA chains, the same ones that alter the behavior of the coding genes;
- Epigenetic Information Layer: Crucial for development, growth, aging, and cancer. It does not alter the DNA sequence although it influences its expression. Epigenetic mechanisms can integrate genomic and environmental signals to control the development of a particular phenotype, and are therefore closely linked to phenotypic plasticity and health.
Epigenetic variations control the activity of genes; if the concentration of substance "X" is high, the activity will be high. The epigenetic code is constituted by a system of molecules linked to DNA or histones, a histone code is the one that governs the expression of genes since their protein tails (those of histones) catalyze a great variety of chemical additions, such as acetyls that amplify neighboring genes.
Epigenetic mechanisms of gene regulation
Epigenetic mechanisms DNA methylation Main article: Methylation It has been discovered that in higher organisms, a methyl group is added to the cytosine base, which allows the closed conformation of chromatin. Therefore a high degree of methylation is associated with gene silencing. One way to control the degree of methylation is through the action of environmental effects. In mammals it has been seen that methionine, choline, folic acid and pyridoxines (which are substances from the diet) have the function of adding methyl groups. In general, methylation occurs to a greater degree in the CpG islands (regions with high concentrations of cytosine and guanine) which are part of the promoter region of genes. For methylation to occur properly, it needs DNA methyltransferase, which is responsible for establishing and maintaining methylation patterns and needs methyl-CpG binding proteins which are involved in making the methylation marks. An example of the importance of silencing a gene or group of genes is the inactivation of the X chromosome and gene imprinting. Gene imprinting refers to the fact that one of the gene copies (it can be both the maternal or paternal copy) that is inherited from the parents, can be completely silenced in order to have a mono-allelic expression of certain genes. Therefore, a methylation pattern corresponding to sex will be observed. If there are abnormalities in the silencing of certain copies, changes in the phenotype may occur that may be the result of diseases such as Beckwith Wiedemann syndrome. This syndrome occurs when the two copies of the IGF2 gene are active, that is, the gene imprinting process did not occur adequately by not silencing the maternal copy, and therefore the individual is characterized by the presence of a high number of large tumors. It has been determined that a high rate of methylation of cell cycle regulatory and DNA repair genes leads to a higher frequency of tumor formation. Similarly, if there is a low level of methylation (hypomethylation), diseases also occur. Recent studies have shown that methylation is a defense mechanism against viruses and parasites to prevent them from damaging DNA.
DNA methylation in cancer DNA methylation is a very important regulator of gene transcription. A very large body of evidence has shown that aberrant DNA methylation is associated with unscheduled gene silencing, and genes that have very high levels of 5-methylcytosine in their promoter region are transcriptionally silenced. DNA methylation is essential in embryonic development, and in somatic cells, DNA methylation patterns are generally transmitted to daughter cells with great fidelity. Aberrant DNA methylation patterns have been associated with a large number of human diseases and have been found in two different ways: hypermethylation and hypomethylation, compared to normal standards. Hypermethylation is one of the major epigenetic modifications responsible for repressing transcription via the promoter region of tumor suppressor genes. Hypermethylation occurs in the CpG islands of the promoter region and is associated with inactivation of the gene. Hypomethylation has also been implicated in the development and progression of cancer through different mechanisms.
Histone modification Chromatin is made up of a basic unit, the nucleosome, made up of histones (H2A, H2B, H3 and H4) bound to non-histone proteins. In the nucleosome the DNA is wound. By post-translational modifications the configuration of histones can be modified. Histones undergo modifications through processes of acetylation, phosphorylation, methylation, deamination, isomerization of prolines, and ubiquitination. Specific combinations in histone modification serve as a kind of code that determines whether the gene is to be silenced or expressed, and this is another way of how gene regulation can occur. Histones.
Non-coding RNA One form of gene regulation is through interference RNAs (iRNAs) which do not code for a specific protein but their sequences are complementary to coding DNA or RNA and prevent their translation, this is a form of negative regulation of expression at the post-transcriptional level. One of these types of RNAs are interference micro RNAs (miRNAs) which bind to complementary sequences and degrade said transcript, thus preventing translation into proteins. The importance of this type of gene regulation has been seen in various scenarios such as: regulation in tumor production, effects of aging due to changes in methylation, associated with stress due to methylation in neural genes, involved in imperfection of fetal development, among others.
All these epigenetic mechanisms play a fundamental role in the correct development and functioning of the organism, as is the case of embryonic development, behavior or cell differentiation, which if uncontrolled can lead to cancer. Epigenetics is responsible for enabling a good organization of chromatin in the cell nucleus, regulating gene expression in different tissues and cell types, and maintaining the correct pattern of expression at the right time and place.
Epigenetics:
A new language DNA is molecularly made up of nucleotides which, in turn, are made up of a sugar, a nitrogenous base and a phosphate. It is precisely the nitrogenous base that distinguishes one nucleotide from another. There are four types of nitrogenous bases in DNA: adenine (A), guanine (G), thymine (T), and cytosine (C). The sequential order of these molecules in the so-called "coding" regions of the genome determines the chemical nature of the proteins that are encoded by these genes and, therefore, their function. In the so-called "regulatory" regions of the genome, the order of the nitrogenous bases defines precisely how the cellular machinery will recognize and process this information. To be functional, DNA molecules must necessarily undergo the process of "transcription" whereby they are faithfully copied to another molecule with a similar chemical nature, RNA, or ribonucleic acid. Regulatory sequences play an important role in this process, that is, sets of nucleotides that are recognized by the cellular machinery or "transcription factors", in such a way that the formation of a multiprotein complex is made possible which, after physically joining the DNA, begins to copy it. The accessibility of regulatory sequences determines the possibility of the transcription process occurring.
Genomic imprinting and epigenetic inheritance Genomic imprinting Methylation processes play an important role in the action of genomic imprinting. In vertebrates this mechanism has only been discovered in mammals. Depending on the parental origin, genes can be activated or silenced. The imprint affects prenatal growth and its importance in the generation of diseases has been established. During gametogenesis, the genomic imprint is initiated and therefore it is inherited during the fusion of the gametes. During the formation of the zygote the imprint is reprogrammed in the new individual. The clearest example of this mechanism occurs in the regulation of the compensatory dose of the X chromosome. This reprogramming plays an important role in the expression of genes in specific tissues that, if they are modified, can have consequences on the proper development of the organism. . Therefore, with a better understanding of how these processes occur and how they are regulated, it is possible to understand diseases such as preeclampsia, losses during pregnancy, failures that occur in assisted reproduction, problems associated with infertility. and cancer among others.
Epigenetic inheritance:
Epigenetic inheritance results from the transmission of information that does not depend on sequences of the nitrogenous bases of DNA through meiosis or mitosis. Epigenetic information therefore modulates gene expression without altering the DNA sequence. DNA methylation patterns are the best studied and understood as markers of epigenetic phenomena.
The epigenome is the global epigenetic information of an organism.
The three main types of epigenetic information are:
- DNA cytosine methylation: it is a change in DNA, in which a methyl group is transferred from S-adenosylmethionine to a C-5 position of cytosine by a DNA-5 methyltransferase. DNA methylation occurs almost exclusively in CpG dinucleotides, having an important role in regulating gene expression.
- Genetic imprint: The imprint is manifested only in higher organisms. When we speak of "imprinting", we refer to genes that can modify their function without the need for a change in the DNA sequence. This change in the way they manifest the "imprinted" genes is generally linked to their parental origin. An imprinted gene manifests in one way when its origin is paternal and in another when it comes from the maternal gamete. It seems that there is a cellular mechanism that somehow "marks" or leaves an imprint on all "printable" genes according to the sex of the individual.
- Histone modification: includes acetylation, methylation, and phosphorylation.
However, when naming these mechanisms, it must be remembered that "indirectly", when analyzing the origin of each process itself, genes are still involved (such as genes for the enzyme DNA-methyltransferase, histones, etc.) and Therefore, genetic changes (such as mutations) that these genes may undergo, or on the genes in which they act, are also indirectly involved. In the same way, although epigenetic modifications do not imply a change in the nucleotide sequence of DNA in the process, but rather consist of a change in the expression of genes, natural selection also, from the biological result of said expression of genes, will act on the epigenetic process and on the organism that manifests it.
Epigenetics in development and phenotypic plasticity
Temperature-dependent effects Enzyme activity depends on temperature, as changes in temperature can affect the way proteins fold, and therefore affect their interaction with other compounds.7 As the phenotype depends on the activity of many Enzymes and their interactions with proteins in general, changes in temperature can result in changes in phenotype.s
Butterflies: changes in the coloration of the wings according to the season Various species of butterflies change their coloration according to the seasons. The changes in coloration have functional advantages, and for this reason they have evolved.7 Usually, the phenotype of the warm summer months has light colors on the wings, while the winter phenotype shows dark colors. Because black colors absorb sunlight more efficiently, they help increase body temperature during winter; the opposite occurs during the summer. The changes in coloration appear to be controlled by the transduction of signals from the environment to the genome through the neuroendocrine system. Environmental signals, such as temperature and length of day, are perceived by the body's neurosensory systems. These can then activate or deactivate the secretion of hormones by the endocrine system. Hormones, in turn, can regulate gene expression, as they can activate transcription factors.
The differences in the coloration of the wings of butterflies in summer and winter are due to changes in the levels of the hormone ecdysone during the larval stage; larvae that develop during cold months are exposed to lower levels of ecdysone than those that develop in warm months.
Reptiles and fish: sex determination Among many species of reptiles, such as turtles and crocodiles, and also in some fish, the sex of an organism depends on the temperature of development of the embryo. This mechanism may have evolved in some species to modify the 1: 1 ratio between the sexes. For example, in crocodiles, high temperatures produce more females, so there can be up to 10 females per male. This represents an advantage for species where the population size is limited by the number of females.
In fish, sex appears to be determined by the relationship between two hormones: estrogen and testosterone, which in turn is controlled by the enzyme aromatase, which converts testosterone to estrogen.7 Temperature can regulate aromatase, and in this way determine the sex of the organism.
Nutrition-dependent effects
Food contains chemical signals that can induce phenotypic changes.
Bees: royal jelly and the queen bee In bees, the production of queen bees depends almost exclusively on feeding the larvae. The larvae that feed on royal jelly (which contains high concentrations of proteins and secretions from the salivary glands of worker bees) throughout their development will be queen bees with functional ovaries. In contrast, larvae that are fed royal jelly for short periods of time will become workers without functional ovaries. Consumption of royal jelly causes high rates of synthesis of juvenile hormone (JH) in the larvae. This hormone delays metamorphosis, allowing the larva to develop longer, acquire a larger size, and develop functional ovaries. The change in JH production levels has been shown to be correlated with the silencing of the Dnmt3 gene, which induces reprogramming of the larval transcriptome. The silencing of this gene is produced by alteration in the levels of methylation, so it is clear that epigenetic regulation is a key component to control the social division of labor in the colony.
Beetles: length of horns The quality and quantity of manure that beetles receive during development determines the morphological and behavioral phenotype of male beetles of some species.15 This occurs in species such as Onthophagus taurus and Onthophagus acuminatus, in which males have horns and females not. As in bees, the juvenile hormone affects the phenotype, this time determining the length of the horns of the males. The higher the juvenile hormone concentrations, the longer the horns.
Feeding-dependent variations in the phenotype of O. taurus males during development As horns are a factor of sexual selection for the female, the behavior of males with short horns changes to ensure their reproduction. While the long-horned males, who have been chosen by the females, guard the door of the den, the short-horned males dig tunnels until they reach the female, to mate with her avoiding confrontation with the long-horned male. . In this way, poor feeding causes low levels of juvenile hormone, which in turn results in short-horned males with "cheating" behavior.
Effects of the presence of predators Some organisms can detect the presence of molecules secreted by their predators and use them to activate the development of structures that make them less susceptible to predation. There are many examples of this ability:
- The cladocero Daphnia produces a head "shaped like a pointed helmet."
- The Keratella rotifer produces additional spines.
- The Chthamalus balano changes the position of its opening.
- The Thais snail produces a thicker shell.
- Tadpoles of the Agalychnis callidryas species begin to hatch from their eggs early in response to vibrations caused by snakes.
Phenotypic variations in Daphnia induced by the presence of predators Effects of the presence of members of the same species Signals to change the phenotype can also come from conspecifics, or members of the same species, since individuals must behave in different ways when they are alone and when they are surrounded by competitors. Usually the signals from predators and conspecifics act synergistically to produce the most favorable phenotype.7 Some examples are shown below:
- Schistocerca gregaria locusts show very different phenotypes with low and high population densities. When the population density is high and resources are scarce, it is beneficial to migrate. The migratory phenotype shows darker colors, longer wings, and aggressive behavior. These changes are caused by odors and direct contact between individuals.
- Fish of many species change sex depending on interaction with conspecifics. For example, in goby fish (Lythrypnus dalli), if the male of the group dies, a female can take his place. But if another larger male is inserted into the group, the converted male can revert his phenotype to female.
The word biomarker refers to any type of variation that occurs in the genetic material and therefore it is possible to detect it in the organism that carries this change. The first markers used are based on the concepts of traditional genetics in such a way that they used polymorphic systems to detect allelic variants that led to a change in phenotype. For the detection of epigenetic modifications, markers were developed that are responsible for detecting molecules that are related to a particular state of activation or inactivation of a gene. For example, the detection of a high amount of methyl molecules indicates a state of inactivation of the gene. With the incorporation of molecular techniques, biomarkers were created through SNPs, indels, RFLPs and microsatellites, among others. For a marker to be considered a good marker, it must require a minimum amount of the sample and must allow the identification of significant differences between a normal state and in a state of epigenetic changes that can develop in a disease. Based on this, there is a type of markers that use body fluids as a sample and measure the concentration of certain metabolites present, which are related to epigenetic changes and later in the formation of cancer. For cancer detection, one of the most frequently used markers are epigenetic modifications of the promoter of genes involved in the inhibition of kinases, dependent on cyclins p15, p16 and RASSF1A. These serve as markers for the early detection of hepatocellular carcinomas. Blood samples are taken from the patient in which methylated sequences of the aforementioned genes can be detected.
Generation of diseases
The knowledge of these phenomena has allowed advances in gene therapies to be made. Work has been done on reversing gene silencing. This work was done in mice with Rett syndrome that, when treated, recovered their ability to produce normal levels of the protein MeCP2, thus reducing the signs of autism that they presented before treatment. A key factor in this field is the heritability of epigenetic marking from one generation to the next, which increases the success of gene therapies. If the structural changes of chromatin can be largely determined by environmental factors and this can be heritable, they would be important in the adaptive expression according to the environment. These latest discoveries have led us to consider not only the expression of genes, but also the way in which said expression can be modified by environmental factors.
Epigenetic regulation is done through structural changes, such as the addition of methyl, which can lead to alterations in the sites of action of enzymes, and as a result, there may be losses in the stability of these regions. Therefore, these regions become more sensitive to chromosomal variations occurring in them or for the cell to transform due to losses in the growth control mechanism or activation of apoptosis. All this can result in changes in the phenotype and a high possibility of the development of diseases.
Epigenetics, Cancer, and NOTCH Signaling Cancer epigenetics is providing new insights into cancer. There are two clear examples of epigenetic modifications in cancer cells: - DNA methylation is believed to be responsible for gene silencing associated with paternal imprinting, heterochromatic gene repression, and X chromosome inactivation. Cancer cells contain DNA methylation patterns modified, that is, they are much less methylated than normal cells. Furthermore, gene promoters in cancer cells are hypermethylated. Therefore, it is believed that these modifications decrease the repression of transcription on most genes that would be silenced in normal cells. These methylation profiles are used today for the diagnosis of tumors. - On the other hand, the modifications in histones, which are also affected in cancer cells.
Since the Notch signaling pathway is known to be involved in both tissue development and renewal, the ideal role played by the NOTCH transduction pathway in cancer proliferation has been raised. Recently, studies have been carried out in Drosophila that have allowed a better understanding of the relationship of NOTCH and tumor formation. NOTCH is important as it has a role in determining cell fate, proliferation, apoptosis, differentiation, migration, and cell development. With the study of drosophila it was determined that the Notch receptors in mammals and the Delta ligands are involved in the formation of tumors. An aberrant activation of the NOTCH1 receptor is related to 50% of types of T-cell acute lymphoblastic leukemia. If the NOTCH transduction pathway is inactivated, tumor formation increases, since it has been seen that in certain contexts Notch can be a tumor suppressor. However, there is still no clear understanding of how Notch acts in cancer formation in vivo. For this reason, studies are focusing on the identification of oncogenes and tumor suppressors that interact with the Notch pathways. With a better understanding of this issue and if it is proven that epigenetic silencing increases tumor formation, epigenetic therapies can be considered to fight cancer.
Cancer epigenetics is an area of ongoing research that continues to facilitate the understanding of the molecular pathogenesis of cancer and the identification of new therapeutic alternatives.19 These investigations have been based on the discovery of biomarkers that facilitate the diagnosis and knowledge of cancer ; DNA methyltransferase and histone deacetylase enzymes have been found to be very effective in cancer diagnosis, and even more specific knowledge about them would aid in the suppression of cancerous tumors.
Possible problems with assisted reproduction It has already been mentioned that many of these changes are the result of exposure to the environment. In the case of assisted reproduction, the question has been raised as to whether the time of exposure to the culture medium may imply an effect on the epigenetic regulation processes. Since the embryo development stage is a critical time in which many epigenetic changes take place (such as a high rate of demethylation to erase the epigenetic marks of the parents), the environment plays an important role to the point that it can change these patterns. During embryogenesis, after the parental imprint is erased, a de novo methylation pattern is formed, which allows tissue differentiation to occur. If this process is not given properly, diseases can occur in the individual or possible problems during pregnancy that can lead to the loss of the embryo. This is why it must be considered that exposure to an artificial environment can have a toxicological potential, preventing the adequate regulation patterns from occurring. Although a large percentage of children born by this technique have normal development and growth, it has been observed that there is a tendency for them to have low birth weight and that there is a three- to six-fold increase in the occurrence of Beckwith-syndromes. Wiedemann and Angelman. There is still no clear knowledge in humans of how the different factors of in vitro fertilization where embryos are formed, can have or have an important effect on their development. Changes in epigenetic imprinting have been observed in mouse embryos.
THE ROLE OF REVERSE TRANSCRIPTASE (RTE)-LIKE AND OTHER NON-LTR RETROTRANSPOSONS IN THE RESPONSE OF SHRIMP TO IHHNV, WSSV, TSV AND IMNV INFECTIONS
|